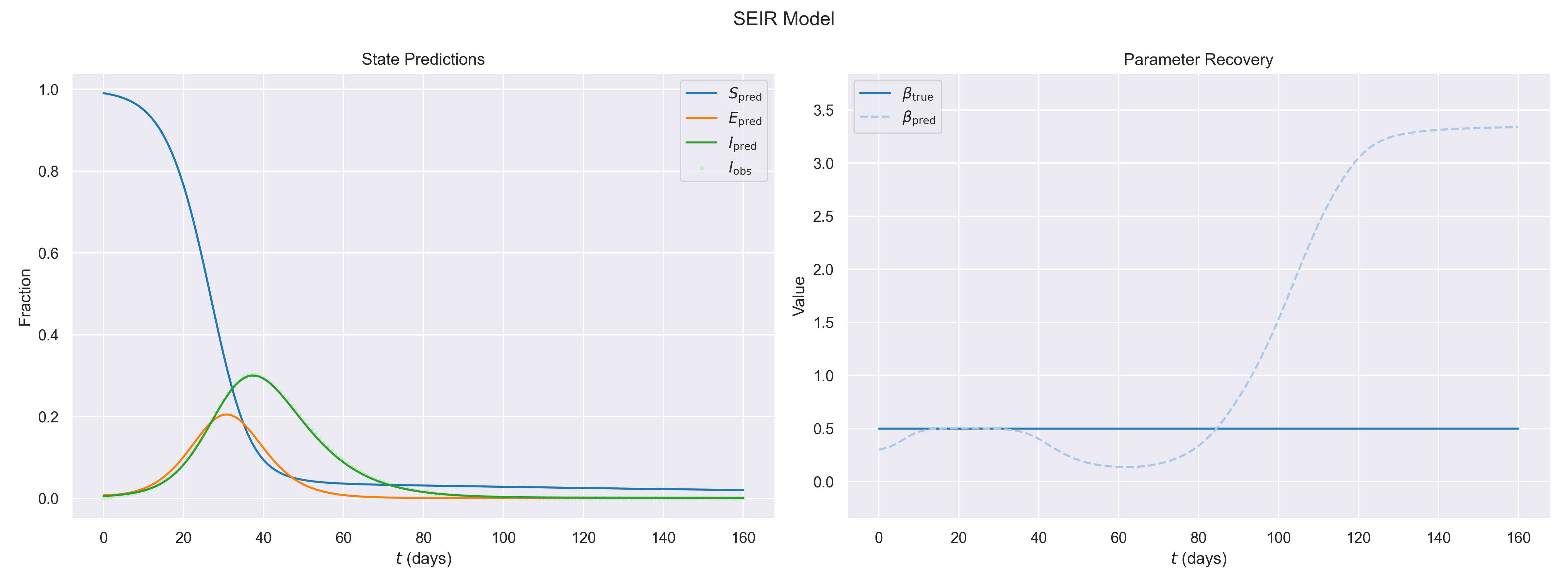
SEIR Epidemic Model
Extended epidemic model with an exposed compartment \(E\). Recovers transmission rate \(\beta\) from partially observed data.
Background
The SEIR model extends the classical SIR framework by introducing an Exposed compartment for individuals who are infected but not yet infectious, capturing the incubation period characteristic of diseases such as COVID-19, measles, and influenza. The latency rate \(\sigma\) (inverse of the mean incubation period) controls the transition from exposed to infectious, adding a delay that significantly affects outbreak timing and peak height.
Governing Equations
The model uses population fractions (\(S + E + I + R = 1\)):
where:
- \(S(t)\): susceptible fraction
- \(E(t)\): exposed (latent) fraction
- \(I(t)\): infected fraction
- \(\beta\): transmission rate (to recover)
- \(\sigma\): latency rate, i.e. \(1 / \text{incubation period}\) (known)
- \(\gamma\): recovery rate, i.e. \(1 / \text{infectious period}\) (known)
The recovered fraction follows by conservation: \(R(t) = 1 - S(t) - E(t) - I(t)\).
Default Configuration
The generated template uses the following values.
Parameters to recover:
| Symbol | Code constant | True value |
|---|---|---|
| \(\beta\) | TRUE_BETA |
\(0.5\) |
Known constants:
| Symbol | Code constant | Value |
|---|---|---|
| \(\sigma\) | TRUE_SIGMA |
\(1/5.2 \approx 0.192\) |
| \(\gamma\) | TRUE_GAMMA |
\(1/10 = 0.1\) |
Initial conditions: \(S(0) = 0.99, \quad E(0) = 0.01, \quad I(0) = 0.001\)
Domain: \(t \in [0, 160]\) days
Features Demonstrated
- Multi-field ODE system (3 fields)
- Scalar
Parameterrecovery ValidationRegistryfor ground-truth comparison
Results