SIR Epidemic Model
Classic S→I→R compartmental model. Recovers transmission rate \(\beta\) from partially observed infected counts.
Background
The SIR model, introduced by Kermack and McKendrick (1927), is the foundation of compartmental epidemiology. It divides a population into three mutually exclusive compartments — Susceptible, Infected, and Recovered, tracking how individuals flow between them. The dynamics are governed by two competing processes: infection at rate \(\beta\) and recovery at rate \(\delta\), whose ratio defines the basic reproduction number \(R_0 = \beta / \delta\). An epidemic occurs when \(R_0 > 1\).
Governing Equations
where:
- \(S(t)\): susceptible population
- \(I(t)\): infected population
- \(N\): total population (constant)
- \(\beta\): transmission rate (to recover)
- \(\delta\): recovery rate (known)
The recovered compartment follows by conservation: \(R(t) = N - S(t) - I(t)\).
Default Configuration
The generated template uses the following values.
Parameters to recover:
| Symbol | Code constant | True value |
|---|---|---|
| \(\beta\) | TRUE_BETA |
\(0.6\) |
Known constants:
| Symbol | Code constant | Value |
|---|---|---|
| \(\delta\) | DELTA |
\(1/5 = 0.2\) |
| \(N\) | N_POP |
\(56 \times 10^6\) |
Initial conditions: \(S(0) = N - 1, \quad I(0) = 1\)
Domain: \(t \in [0, 90]\) days
Scaling: populations are divided by \(C = 10^6\) and time is normalized by \(T = 90\) in the training ODE. The generated code maps between physical and scaled units automatically.
Features Demonstrated
- Scalar
Parameterrecovery ValidationRegistryfor ground-truth comparisonDataScalingcallback for population-scale normalization
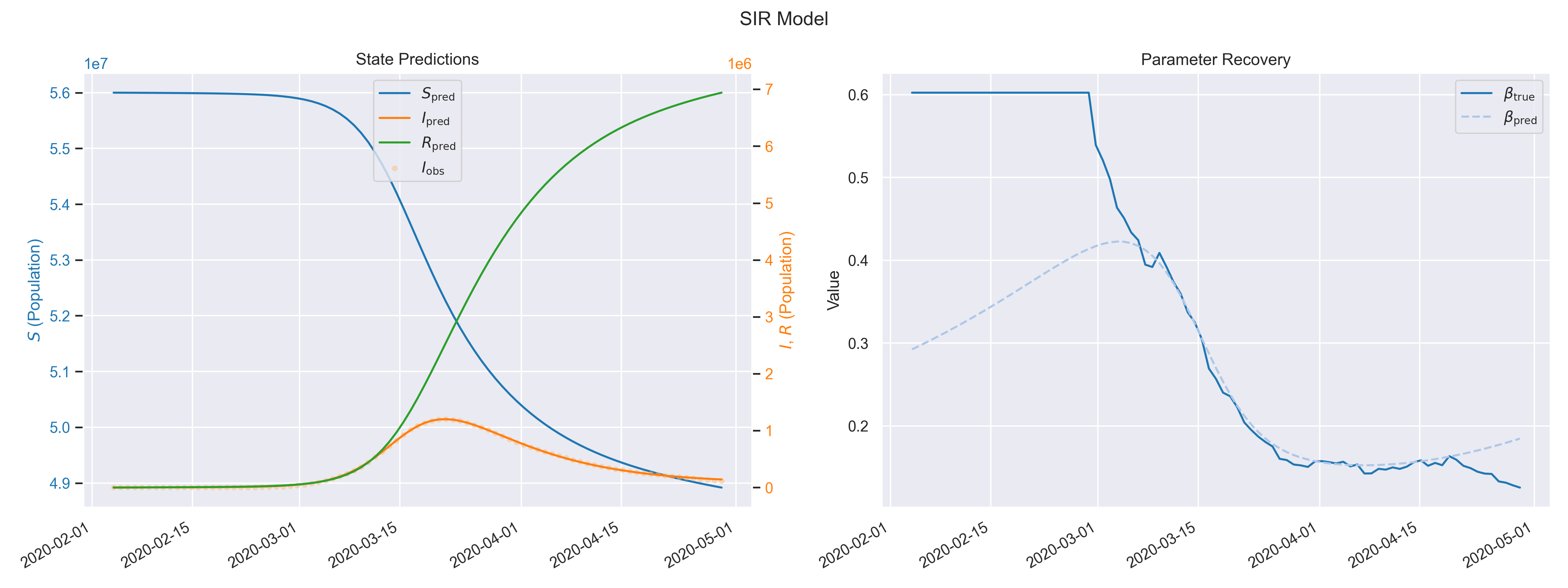
Results